Over 90% of Microbiome Experiments Show Disease—Scientists Say That’s Implausible
Systematic reviews reveal a near-perfect success rate — raising concerns about bias, pseudoreplication, and how microbiome research is done.
Animal models are supposed to test our hypotheses, not mirror our expectations. But when scientists reviewed animal studies on gut microbiome transfer, they noted something that’s statistically suspicious: almost every experiment reported successful transmission of disease.
If true, this would mean all diseases do indeed begin in the gut. If that sounds too simplistic to be true to you, your gut is probably right. Because it’s out of proportion to the biological complexity in humans.
These scientists were Professor Jens Walker and colleagues at leading Irish and Canadian institutions, who argued that the 95% success rate seen in animal models of gut microbiome transferring diseases is biologically implausible. They published their work in the prestigious Cell journal in 2019, titled “Establishing or Exaggerating Causality for the Gut Microbiome: Lessons from Human Microbiota-Associated Rodents.”
I just came across this 2019 paper while researching and writing about problems with the gut microbiome research the other day. So, this article also serves as a supplementary piece to my recent piece, “The Microbiome Was Supposed to Change Medicine. What Happened?”
Although I’m 7 years late, things don’t seem to have changed much, as we’ll explore later. But that’s not surprising. Seven years is barely a generation of research; it’s roughly the length of a PhD in the U.S.
I. The 95% Success Rate
Walker et al. did a systematic review of human microbiota-associated (HMA) rodent studies. These kinds of animal studies transplant the gut microbiome, typically from humans with a disease, into rodents raised in a sterile environment. So, these rodents are germ-free, lacking a microbiome, making them highly permissive to gut microbiome transplantation.
(Note: Systematic review screens studies across databases with predefined keywords and eligibility criteria. This minimizes bias seen with typical literature reviews, which are prone to cherry picking, i.e., reviewing studies that fit a hypothesis and ignoring studies that don’t.)
Such a study design makes causal (cause-and-effect) inferences powerful. After all, if a rodent gets the same disease humans do after receiving the human gut microbiome, that would mean the gut microbiome is the source of the disease and that the disease indeed began in the gut.
Scientists have successfully transferred various diseases from humans to rodents through this method, including obesity, asthma, allergies, inflammatory bowel disease, colorectal cancer, Parkinson’s disease, Alzheimer’s disease, depression, anxiety, and autism.
A systematic review of these studies by Walker et al. (Figure 1) revealed that, out of 38 eligible studies that met their criteria, 36 reported that transferring gut microbiome from diseased humans into germ-free rodents led to at least one disease-related trait appearing in the animals.
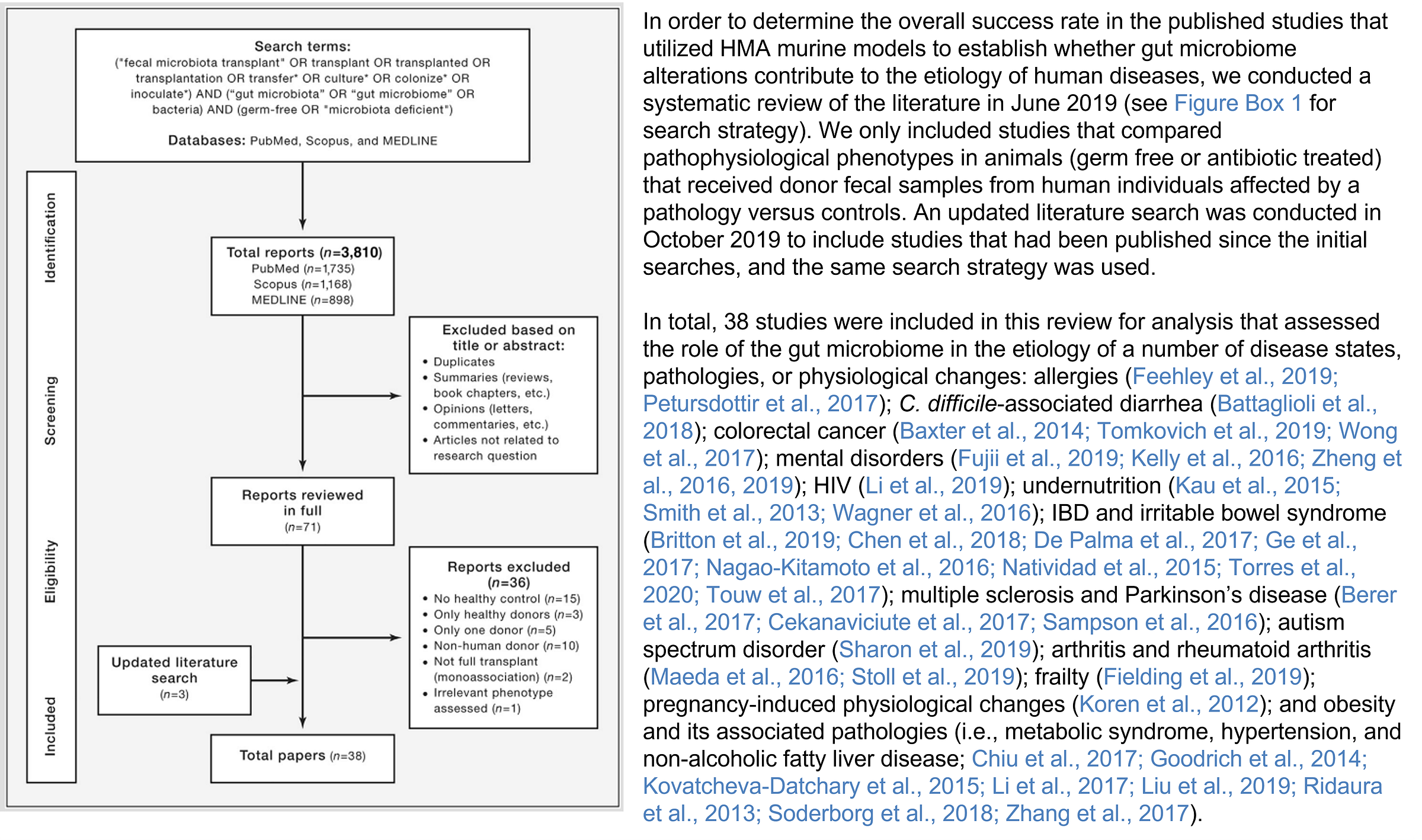
That’s 95% success rate.
So, almost every time the gut microbiomes from sick people were transferred into sterile rodents, the rodents developed features of that disease—be it inflammation, altered behavior, or immune changes—compared with rodents that received microbiomes from healthy humans.
What about the two studies that comprise the 5% failure rate?
According to Walker et al., these two exceptions were studies on colorectal cancer. But even these exceptions didn’t show that the microbiome had no effect at all. They simply showed that the outcome wasn’t strictly determined by whether the human donor had cancer.
Instead, the cancer in mice depended on other microbial factors—such as the mouse’s existing microbiome or microbes from colon biofilms—thus still implicating the gut microbiome in disease to some extent.
So, the systematic review concluded that:
36 out of 38 (95%) were positive,
2 out of 38 (5%) were “complicated,”
and none truly contradicted microbiome causality.
That level of uniformity is what Walker et al. found suspicious.
They argued that in complex biological systems, especially across species, such near-universal confirmation is unusual. Human diseases like obesity, depression, or neurodegenerative disease are shaped by genetics, environment, and lifestyle, not single-variable problems.
So how could transferring gut microbes into mice reproduce disease traits almost every time?
II. The Methodological Problem
Walker and colleagues argue that part of the answer may be how these experiments were designed and analyzed.
The most important issue is something called pseudoreplication.
To understand this, we need to clarify what counts as a true sample. In microbiome transfer experiments, the real unit of interest is the human donor. Researchers are asking whether microbiome from different people—sick versus healthy—produce different outcomes in animals. So the true sample should be the number of human donors included.
But in many studies, only a small number of human donors were used. Their microbiome was then transplanted into multiple mice. Those mice were treated as independent samples in the statistical analysis.
On paper, this inflates the sample size. Ten or twenty mice per group may look statistically powerful. But if those mice all received microbiota from the same one or two human donors, they are not truly independent biological replicates. They are repeating the same donor signal.
Walker et al. found that 84% of the studies used the individual animals as the statistical N, even when the number of human donors was much smaller. In essence, many studies have exaggerated their statistical confidence by counting mice rather than human donors as their sample.
As the human gut microbiome varies enormously between individuals, using only a handful of donors does not capture that diversity. A small, unrepresentative group of donors may accidentally produce consistent results — even if the broader population would not.
This pseudoreplication problem gets worse when studies pool donor samples. Some studies combined samples from multiple donors into one mixture before transplanting them into mice. While this may seem practical, it further reduces the number of true experimental units. If five donors are pooled into a single mixture, the study has only one donor sample, not five.
Walker et al. also pointed to another weakness: many studies did not confirm that the gut microbiome imbalance (dysbiosis) observed in humans was reproduced in mice. Specifically:
Only 63% (24/38) of studies confirmed the presence of dysbiosis in the original human donor samples.
Only 29% (11/38) confirmed that some aspect of that dysbiosis—such as reduced diversity, shifts in bacterial groups, or altered metabolic capacity—was successfully transferred into the recipient animals.
So, studies often showed that disease traits appeared in mice, but did not always verify that the human microbial imbalance itself exists and, if it does, has successfully transferred into the mice.
Another concern Walker et al. raised was that most studies stopped at showing an effect, without explaining what actually caused it.
Out of the 38 studies reviewed, only 13 (34%) attempted to explore underlying mechanisms linking the altered microbiome to disease. The remaining 66% of studies did not delve deeper to identify which specific microbes were responsible or which biological pathways were being altered.
Walker et al. argue that this mechanistic depth is one of the major advantages of using animal models, which allow researchers to isolate individual microbes, track gene changes, and test biological pathways directly. Yet most of the studies claiming causality did not take this extra step.
Overall, these methodological problems—pseudoreplication (small numbers of human donors and sample pooling) and limited mechanistic investigation (not confirming dysbiosis and lack of molecular investigation)—exaggerate the positive findings (Figure 2).
This doesn’t mean the results were fabricated. Rather, it suggests that the experimental design may bias outcomes toward detecting a positive effect. And when nearly every study detects an effect, even across wildly different diseases, it becomes reasonable to ask whether the model is mirroring our expectations on causality rather than testing it.
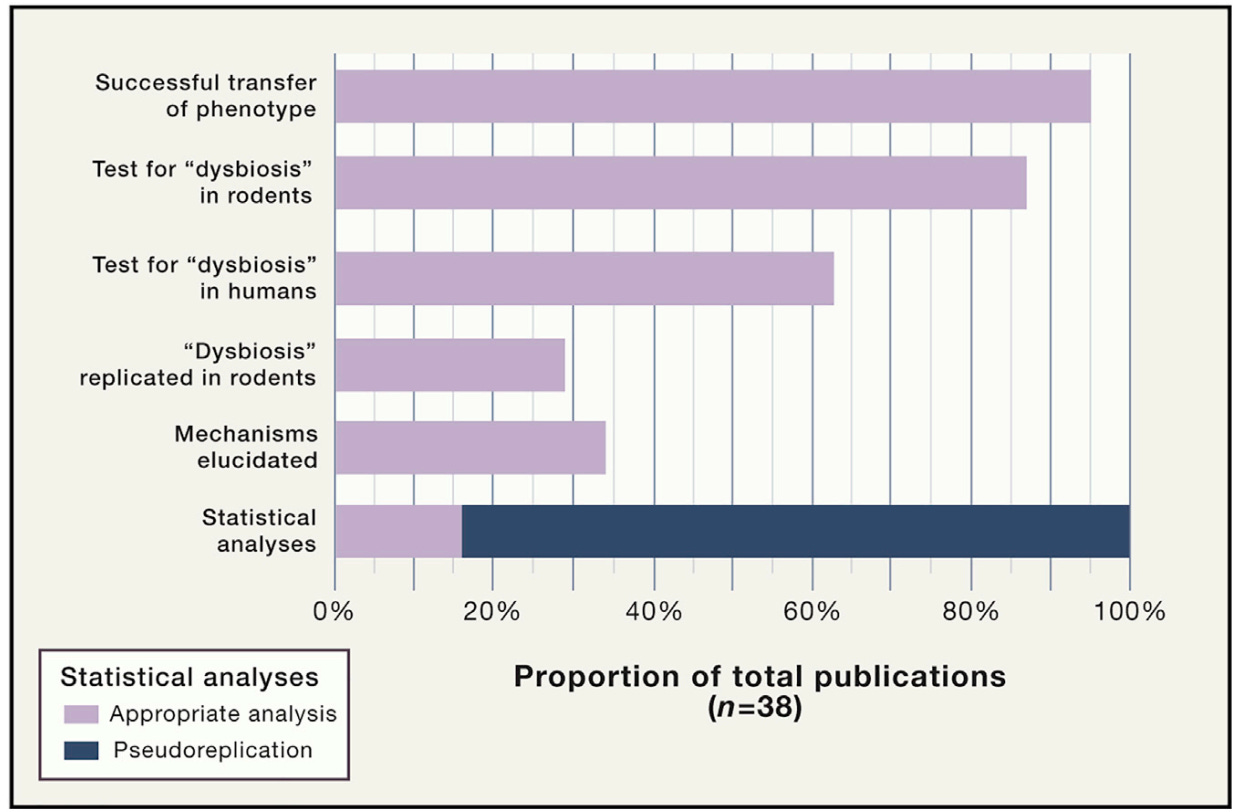
III. The Biological Problem
There is another reason the 95% success rate is difficult to accept at face value: biology does not transfer easily across species.
Even well-established human gastrointestinal pathogens often fail to reproduce the same disease pathology in mice without drastic experimental manipulation:
Helicobacter pylori: A confirmed cause of gastritis, ulcers, and stomach cancer in humans. Yet it does not colonize the gastrointestinal tract of mice in a way that reproduces the human disease.
Clostridioides difficile: A well-known cause of antibiotic-associated colitis in humans. In standard mouse models, it does not consistently establish infection or replicate human-like pathology.
Enteropathogenic and enterohemorrhagic Escherichia coli: These strains cause severe intestinal disease in humans, but they do not naturally produce disease in mice. Instead, mice develop illness from a related mouse-adapted pathogen, Citrobacter rodentium.
Salmonella enterica serovar Typhimurium: In humans, this bacterium typically causes limited intestinal diarrhea. But in mice, it produces a systemic infection involving the liver and spleen — a disease pattern more similar to typhoid fever than to typical human salmonellosis.
Campylobacter jejuni, Vibrio cholerae, and Shigella species: A common cause of bacterial gastroenteritis, cholera, and dysentery in humans, respectively. But these pathogens do not reliably colonize or reproduce human-like disease in standard mouse models.
So, even single, well-defined human gastrointestinal pathogens struggle to replicate their full disease behavior across species.
If that is true for individual pathogens with specific virulence factors, it raises an obvious question: How likely is it that an entire human microbial community — containing thousands of interacting species — can consistently recreate complex, multifactorial human diseases in mice?
Walker et al. argue that these cross-species barriers make the near-perfect success rate of microbiome transfer studies even more implausible.
A second biological reason not discussed in the Walker et al. paper is that germ-free mice are inherently designed for success.
These animals are raised in sterile environments without any microbes. Because their immune systems and gut ecosystems develop without exposure to microbes, these mice are very responsive to microbial exposure later in life. Introducing a new microbial community into such an empty ecological space can produce large biological effects.
Moreover, laboratory animals are genetically similar, housed in standardized environments, and fed identical diets. These tightly controlled conditions reduce many of the variables that influence health and disease.
But human populations look very different. People vary widely in their genetics, diets, environments, lifestyles, and microbiomes. Their immune systems and microbial ecosystems have also developed over many years of exposure to different microbes and environmental factors.
IV. The Sociology of Academics
Walker and colleagues raised another factor contributing to the near-perfect success rate of gut microbiome transfer studies: positive-result bias.
In academic publishing, studies with clear positive findings are far more likely to appear in high-impact journals. Negative results — studies that fail to confirm a hypothesis — are much harder to publish and often end up in less prestigious journals or remain unpublished. This creates what researchers call the “file drawer problem” or “publication bias,” where studies that do not support a hypothesis quietly disappear from the scientific record.
As a result, the published literature can give a distorted impression of how often experiments actually succeed. If negative studies are missing, systematic reviews will naturally be dominated by positive reports.
As Walker and colleagues wrote in their paper, “Consequently, investigators (this includes the authors of this review) are incentivized to withhold data that do not support their hypotheses or to recast it in a way where negative findings are presented as positive results.”
Walker et al. also noted that this phenomenon is not unique to microbiome science. Positive-result bias is widely recognized across biomedical research. Researchers, institutions, journals, and even the media all tend to favor promising findings over ambiguous or null results (Figure 3).
That said, this issue may be particularly important for microbiome transfer studies. These experiments sit at the center of a major scientific question: whether changes in the gut microbiome can “cause” human disease. Because the results speak directly to causality, positive findings are especially attractive to academic journals, media, and readers.
But scientific progress relies on self-correction — the ability of new findings to confirm, refine, or contradict earlier ones. That process only works when negative results are published as well. If not, the literature risks reinforcing conclusions that may be less certain than they appear.

V. The Pattern Continues
Ever since Walker et al.'s paper was published in 2019, the broader literature suggests the field has not fundamentally corrected course.
A 2021 systematic review examining 241 gut microbiome transfer studies still found that 92.5% reported a positive outcome. When the authors looked closer, they found that many papers failed to report basic experimental details, such as sample preparation, storage conditions, microbial concentration, administered volume, and delivery route.
These methodological gaps were serious enough to make replication difficult, prompting the authors from renowned Australian and Irish universities to propose the new Guidelines for Reporting Animal Fecal Transplantation (GRAFT) to improve transparency and reproducibility.
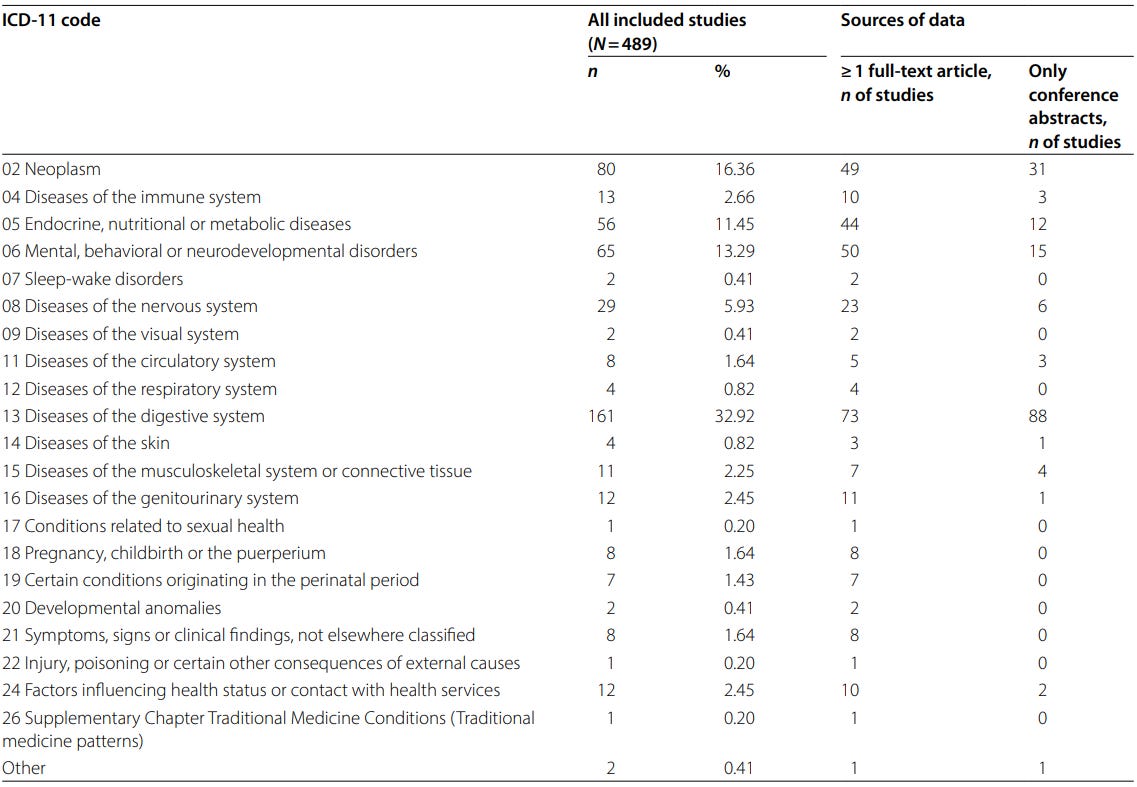
A larger 2025 scoping review of 489 microbiome transfer studies reached a similar conclusion. The analysis showed that these experiments have been applied across a wide range of diseases, including cancer, metabolic disorders, neurological, and psychiatric conditions (Figure 4).
Despite this diversity, these experiments report a remarkably high success rate of more than 80%. But the authors noted that experimental protocols also vary widely. Differences in donor selection, sample preparation, antibiotic pretreatment, housing conditions, and outcome measures make it difficult to compare results across studies or to determine causality.
This means it’s unclear whether the microbiome is causing the disease or whether the experimental setup is producing the effect.
Even in more specialized areas such as the gut–brain axis, the same problem persists. In a 2026 systematic review of 31 studies that transferred microbiomes from psychiatric patients into rodents, there was still high methodological variation across studies. As many as 18 studies pooled fecal samples, and 15 studies did not check whether the human microbiome had actually engrafted in the recipient animals
So, even more recent literature suggests that many of the concerns Walker et al. raised — unusually high success rates, lack of method standardization, pooling of donors, and incomplete confirmation that microbiome transfer actually occurred — still remain prevalent.
VI. Rebuilding the Evidence
None of these criticisms means that human microbiota-associated (HMA) animal models should be abandoned. Walker et al. are careful to emphasize that these models remain valuable tools for studying host–microbe interactions. But if they are going to be used to support strong claims about human disease, the standards need to be much higher.
That starts with better experimental design. Studies need enough human donors to reflect real biological variation, and they need to stop treating mice as independent samples. Donor samples should not be pooled in ways that erase individual variability, and researchers should confirm that the microbial patterns seen in humans actually engraft in recipient animals.
Just as importantly, studies should move beyond simply showing that a disease-related trait appears in a mouse. The real value of these models lies in identifying mechanisms — the specific microbes, pathways, metabolites, or ecosystem functions that may be driving the effect.
But Walker et al. also argue that causality cannot be settled in mice alone. Because findings in mice do not always translate to humans, newer approaches are emerging to test causality in people. These include causal inference tools such as Mendelian randomization, as well as longitudinal studies that track whether microbiome changes appear before or after disease. If microbiome changes consistently precede worsening of a disease marker, that is more informative than observing both microbiome changes and disease at the same time (chicken-or-egg problem).
Perhaps the most convincing approach is an interventionist framework. The idea is simple: if you change a suspected cause and the disease process changes, that provides strong causal evidence. This logic helped establish Helicobacter pylori as a cause of peptic ulcer disease and the microbiome as a protective factor against C. difficile infection. Applied to the microbiome, this could mean deliberately altering microbial communities, specific species, or even metabolites, then observing whether host outcomes change.
In the end, Walker et al. are calling for a shift in mindset. The goal of microbiome science should not be to link the gut to every disease imaginable. It should be to figure out which microbial changes truly matter, how they work, and whether changing them actually does anything.
If you’ve made it this far, thank you for reading! Most of my Substack articles are paywalled, so if you found this valuable, consider subscribing for just $2.90/month (annual plan). Your support allows me to devote more time to researching and writing about lesser-known but crucial topics like this.




Good science practiced here! Thank you so much.